Tetra File Converter
Package your heterogenous experiment data sets into one single file, leveraging Allotrope Ontology and HDF5, designed for data science and cloud computing
Overview
ADF is short for Allotrope Data Format. Tetra File Converter converts an IDS JSON into an ADF file. To understand the technical content on this page, we recommend you first read the following:
- Allotrope Leaf Node Model — a Balance between Practical Solution and Semantics Compatibility
- Tetra ADF Converter
- Intermediate Data Schema (IDS) JSON
Design Philosophy
Tetra ADF Converter operates with the following design philosophies.
- Simplification, while preserving semantic extensibility. Use Leaf Node Model to make automation of the RDF construction possible while ensuring that Full RDF Graphs can be built on top of the Leaf Node RDF TTL files.
- Data Science ready. The resulting files can be opened and read using any tools that support HDF5 and software developers and Data Scientists can understand the content of the ADF file without needing to learn semantic web or RDF.
- Automated. Use Web API to ingest IDS JSON to be converted and use Web API to retrieve the results.
Conversion Logic
Input (IDS JSON)
{
"@idsType": "example",
"@idsVersion": "v1.0.0",
"@idsNamespace": "common",
"system": {
"serial_number": "serial_number"
},
"time": {
"measurement": "2015-09-24T03:47:13.0Z"
},
"sample": {
"id": "unknown-10",
"batch": {
"id": "batch-number"
}
},
"method": {
"instrument": {
"cell_type": "CHO",
"dilution_factor": 1
}
},
"user": {
"name": "operator-1"
},
"result": {
"cell": {
"viability": {
"value": 0.1,
"unit": "Percent"
},
"diameter": {
"average": {
"live": {
"value": 21.07,
"unit": "Micrometer"
}
}
},
"count": {
"total": {
"value": 1207,
"unit": "Cell"
},
"viable": {
"value": 1,
"unit": "Cell"
}
},
"density": {
"total": {
"value": 102.24,
"unit": "MillionCellsPerMilliliter"
},
"viable": {
"value": 0.1,
"unit": "MillionCellsPerMilliliter"
}
}
}
},
"peaks": [{
"area": {
"value": 111.1,
"unit": "AU"
},
"retention_time": {
"value": 111.1,
"unit": "Minute"
}
}, {
"area": {
"value": 222.2,
"unit": "AU"
},
"retention_time": {
"value": 222.2,
"unit": "Minute"
}
}],
"datacubes": [{
"name": "3D chromatogram",
"description": "More information about the data cube. (Optional)",
"measures": [{
"name": "intensity",
"unit": "ArbitraryUnit",
"value": [
[111, 222, 333, 444, 555],
[111, 222, 333, 444, 555],
[111, 222, 333, 444, 555]
]
}],
"dimensions": [{
"name": "wavelength",
"unit": "Nanometer",
"scale": [180, 190, 200]
}, {
"name": "time",
"unit": "MinuteTime",
"scale": [1, 2, 3, 4, 5]
}]
}]
}Data Description
- ADF Converter will traverse the entire IDS JSON and identify all the Leaf Nodes and then generate the corresponding triples to form the TTL file leveraging the ontology IRI and prefLabels embedded in the IDS schema file.
- The generated triples follow the Allotrope Leaf Node pattern.
- The TTL file will be inserted into the ADF file's Data Description section using Allotrope JAVA Library.
IDS JSON
{
"sample": {
"id": "abc"
}
}IDS Schema.json
{
"type": "object",
"properties": {
"sample": {
"type": "object",
"properties": {
"id": {
"type": "string",
"@type": "http://purl.allotrope.org/ontologies/result#AFR_0001118",
"@prefLabel": "sample id",
"description": "this is the id or barcode of your sample"
}
}
}
}
}Resulting TTL
<urn:uuid:94b87bf1-245f-4487-ad93-5539b0c544c9>
a <http://purl.allotrope.org/ontologies/result#AFR_0001118> ;
<http://purl.allotrope.org/ontologies/property#AFX_0000690>
"abc"^^<http://www.w3.org/2001/XMLSchema#string> .
<http://purl.allotrope.org/ontologies/result#AFR_0001118>
<http://www.w3.org/2004/02/skos/core#prefLabel>
"sample id" .Data Cubes
- Data Cubes from the IDS JSON will be inserted into the ADF file's Data Cubes section using the Allotrope JAVA Library
Recommended For the Data ScientistsAllotrope JAVA Library has limitations that it does not attach human-readable units to the resulting HDF5 data sets and also the measure and dimensions do not have human-readable names.
For users who do not want to understand/use SPARQL queries, Semantic Web or RDF to just make sense of the Data Cubes, Tetra File Converter provides a configurable option for you to use HDF5 Python library h5py to create the Data Cubes. If you choose to use this approach, Tetra File Converter will attach the unit as data set attributes and also use human-readable labels for the measures and dimensions.
To enable this option, simply attach "hdf5: true" as custom metadata to the IDS JSON you ingest.
Data Packages
- The TTL file, IDS JSON file will both be inserted into the ADF file's Data Package section using Allotrope JAVA Library.
- All the files referenced via file pointers in the IDS JSON will be inserted into the ADF file's Data Package section using Allotrope JAVA Library.
Output
- The conversion will generate ADF file or or HDF5 file, depending on which approach is used.
- The conversion will also generate related artifacts:
- Data description TTL
- Leaf Node JSON
FAQ
Where are the ADF files stored ? How can I search find and use the ADF files?
All the ADF files will be stored in the Tetra Data Lake inside the Tetra Data Platform. We provide web RESTful API to search and retrieve the files. For example, find me all ADF files for a particular type of instrument and created in the last two days. Search API can be found at Search files via Elasticsearch query language and retrieved via Retrieve a file API.
How can I move the ADF file to another system?
You can configure a Data Pipeline to push ADF files or the content of the ADF files to a targeting system.
How can we convert data from RAW instrument files to ADF or HDF5?
You may combine this application with the Tetra Data Platform, which will acquire data from the instruments, move the data to the cloud and then harmonize the data into IDS JSON.
Why inserting the TTL and IDS files into the ADF Data Package?
Currently, in order to read the Data Description section of the ADF files, you HAVE TO use the Allotrope JAVA and .NET library and learn SPARQL query and RDF. See below, the ADF file is opened via the HDF5 Viewer, the Data Description section is not readable using common HDF5 tools.
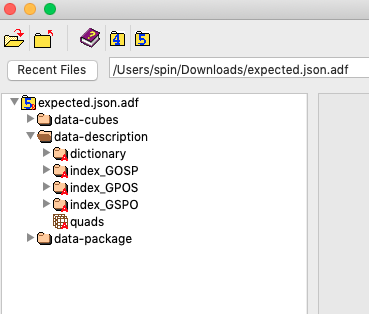
This greatly limits Data Scientists' ability to consume the ADF files using common tools like Python, R, Jupyter Notebook.
Tetra File Converter puts the TTL and IDS JSON into the Data Package, thus making it possible for Data Scientists and Software Developers to
- Read the content of the ADF file without learning RDF or SPARQL (just use the IDS JSON).
- Read the Data Description RDF TTL using any HDF5 compatible library, such as h5py (just read the TTL file)
Updated 2 months ago
